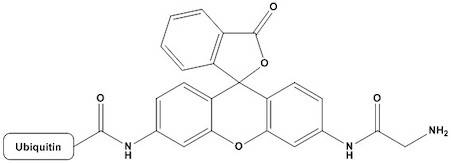
| Molecular Formula: | C400H644N108O121S |
| Molecular Weight: | 8933.9 |
| Ubiquitin-Rhodamine 110 (Ub-Rho110) is a fluorogenic substrate widely used to study deubiquitylating enzymes (DUBs) and ubiquitin-like processing enzymes. It consists of ubiquitin — a highly conserved 76-amino acid protein — fused at its C-terminus via an amide bond to Rhodamine 110 (Rho110), a fluorescent dye. In its intact form, the molecule exhibits minimal fluorescence due to quenching of the rhodamine moiety by the attached ubiquitin peptide. Upon cleavage of the amide bond by a DUB or ubiquitin C-terminal hydrolase (UCH), free Rhodamine 110 is released and produces a strong, easily measurable fluorescent signal. This elegant design makes Ub-Rho110 one of the most widely adopted tools in ubiquitin research, particularly for enzyme activity assays, high-throughput screening (HTS) campaigns, and mechanistic studies. Since its introduction, the substrate has enabled discoveries in cancer biology, neurodegenerative disease research, immunology, and the development of novel DUB-targeting therapeutics. 1. Biochemical Basis and Mechanism of Action 1.1 Ubiquitin Biology Ubiquitin is a small regulatory protein found in virtually all eukaryotic cells. The ubiquitin system governs a broad array of cellular processes including protein degradation via the 26S proteasome, DNA damage response, cell cycle regulation, endocytosis, and immune signaling. Ubiquitination involves the sequential action of three enzyme classes: E1 (ubiquitin-activating), E2 (ubiquitin-conjugating), and E3 (ubiquitin ligase) enzymes. These enzymes attach ubiquitin to substrate proteins via an isopeptide bond between the C-terminal glycine (Gly76) of ubiquitin and a lysine residue on the target. Reversal of ubiquitination is catalyzed by DUBs, which comprise approximately 100 enzymes in the human genome. DUBs are cysteine or metalloprotease enzymes that cleave ubiquitin from substrates or disassemble polyubiquitin chains. Their critical role in cellular homeostasis has made them attractive therapeutic targets. 1.2 Rhodamine 110 as a Fluorogenic Reporter Rhodamine 110 is a xanthene-class fluorescent dye with excitation and emission maxima at approximately 496 nm and 520 nm, respectively. It is one of the most sensitive fluorophores available, with a high molar extinction coefficient (~80,000 M⁻¹cm⁻¹) and quantum yield (>0.9 in aqueous solution). Critically, the free amine groups of Rhodamine 110 are required for full fluorescence; acylation or amide bond formation at these positions dramatically reduces fluorescence intensity due to disruption of the conjugated electronic system. In Ub-Rho110, the C-terminus of ubiquitin is coupled to the amine of Rhodamine 110 via an amide bond. This bond effectively quenches the fluorophore. Hydrolysis of this bond by a DUB restores the free amine and generates a large increase in fluorescence signal — typically a 100- to 1000-fold enhancement over background — providing an extremely sensitive readout of enzymatic activity. 1.3 Kinetics of Fluorogenic Signal Development The fluorescence signal generated by DUB cleavage of Ub-Rho110 follows standard Michaelis-Menten kinetics. The assay can be run in two modes: continuous (kinetic), in which fluorescence is monitored in real time, or endpoint, in which the reaction is stopped and fluorescence is measured at a fixed time point. Continuous monitoring is preferred for determining kinetic parameters such as kcat, KM, and kcat/KM, while endpoint assays are amenable to HTS workflows using microplate readers. Excitation maximum: 496 nm (Free Rhodamine 110) Emission maximum: 520 nm (Free Rhodamine 110) Molar extinction coefficient: ~80,000 M-1cm-1 (At 496 nm) Quantum yield: >0.9 (In aqueous buffer) Fluorescence enhancement: 100– >1000-fold (Upon DUB cleavage) Typical assay concentration: 0.1–10 μM (DUB-dependent) 2. Physical and Chemical Properties 2.1 Molecular Structure Ub-Rho110 consists of full-length human ubiquitin (residues 1–76) with its C-terminal carboxylate (Gly76) coupled via a stable amide bond to the 6-amino group of Rhodamine 110. The resulting conjugate has a molecular weight of approximately 9.7 kDa (combining ubiquitin at ~8.6 kDa and Rhodamine 110 at ~331 Da). The compound is soluble in aqueous buffers and commonly supplied as a lyophilized powder. 2.2 Solubility and Storage Ub-Rho110 is typically reconstituted in aqueous buffer at neutral to slightly alkaline pH (e.g., 50 mM HEPES pH 7.5, or PBS pH 7.4) supplemented with a reducing agent such as 1–2 mM dithiothreitol (DTT) or tris(2-carboxyethyl)phosphine (TCEP). Stock solutions are generally prepared at 100–500 μM and stored at −80°C in single-use aliquots to avoid repeated freeze-thaw cycles, which can lead to partial hydrolysis and an increase in background fluorescence. Lyophilized product is stable for at least 12 months at −20°C to −80°C when stored in the dark. 2.3 Stability Considerations The amide bond in Ub-Rho110 is chemically stable under the recommended storage conditions, but non-enzymatic hydrolysis can occur under acidic or strongly alkaline conditions, as well as at elevated temperatures. Importantly, background fluorescence must be assessed in every assay by including enzyme-free controls. Exposure to light should be minimized throughout handling, reconstitution, and assay setup, as rhodamine dyes are susceptible to photobleaching. 3. Applications 3.1 Measurement of DUB Activity The primary and most extensively validated application of Ub-Rho110 is as a fluorogenic substrate for DUB activity assays. Virtually all cysteine DUBs (UCH, USP, OTU, MINDY, ZUFSP families) and JAMM/MPN metalloprotease DUBs can cleave the Ub-Rho110 amide bond, making this a broadly applicable substrate. Some notable examples include: • UCH-L1, UCH-L3, and UCH-L5: Among the most active DUBs toward Ub-Rho110, with high catalytic efficiency (kcat/KM ~105–106 M⁻¹s⁻¹). • USP2, USP5, USP7, USP14, and USP21: Well-characterized USP family members routinely tested against Ub-Rho110 in substrate profiling studies. • OTULIN and OTUD family enzymes: OTU domain DUBs with varying efficiency toward Ub-Rho110 depending on the specific linkage preference. • AMSH and AMSH-LP: JAMM metalloprotease DUBs that can cleave Ub-Rho110, though metalloprotease DUBs generally show lower activity toward this substrate than cysteine DUBs. The assay typically requires the DUB enzyme to be activated, which for cysteine DUBs involves pre-incubation with a reducing agent (DTT or TCEP) to ensure the active-site cysteine is in the reduced, nucleophilic state. 3.2 Enzyme Kinetics and Mechanistic Studies Ub-Rho110 is widely used to determine kinetic parameters of DUBs under varying substrate concentrations. The continuous fluorescence readout allows real-time monitoring of product formation, enabling precise measurement of initial velocities (V0) as a function of substrate concentration. Lineweaver-Burk and Michaelis-Menten analyses with Ub-Rho110 have provided kinetic constants for dozens of DUBs and have been instrumental in understanding catalytic mechanisms, including the role of active-site residues, conformational changes upon ubiquitin binding, and the impact of post-translational modifications on DUB activity. The substrate is also used in pH-rate profile studies, temperature dependence analyses, and solvent kinetic isotope effect experiments to probe the chemical mechanism of DUB-catalyzed reactions. 3.3 High-Throughput Screening for DUB Inhibitors The high signal-to-noise ratio, simple assay format, and compatibility with standard microplate readers make Ub-Rho110 particularly well suited for HTS campaigns aimed at identifying small-molecule DUB inhibitors. The assay has been miniaturized to 384- and 1536-well formats and is compatible with automated liquid handling systems. Typical assay validation metrics include: Z’ factor : >0.6 (≥0.5 considered robust) Signal-to-background ratio: 5– >100 (≥5 preferred) Coefficient of variation (CV): <10% (<15% acceptable) DMSO tolerance: 1–2% (Enzyme-dependent) Assay volume (384-well): 10–20 μL (Optimized per target) Several DUB inhibitors now in preclinical or clinical development were first identified in HTS campaigns using Ub-Rho110, highlighting the substrate’s translational value. 3.4 Profiling DUB Activity in Cell Lysates Ub-Rho110 can be used to measure total DUB activity in crude cell or tissue lysates. This approach has been employed to compare DUB activity between normal and diseased tissues, to assess the impact of genetic knockdown or overexpression of specific DUBs, and to evaluate pharmacodynamic responses to DUB inhibitors in cell-based studies. However, interpretation of lysate-based assays requires caution, as multiple DUBs contribute to total signal and the relative contributions may vary depending on the sample source and preparation method. 3.5 Activity-Based Protein Profiling (ABPP) Although Ub-Rho110 itself is not an activity-based probe (ABP), it is often used alongside ABPs in DUB profiling studies. The combination of fluorogenic substrate assays (to measure total enzymatic activity) with ABP-based chemoproteomics (to identify and quantify specific DUBs) provides a comprehensive picture of DUB activity in biological systems. Ub-Rho110 is also frequently used to confirm the activity of recombinant DUBs before their use in ABPP experiments. 3.6 Structural and Biophysical Studies The well-defined structure of ubiquitin and the availability of high-resolution crystal structures of ubiquitin-DUB complexes have enabled the rational design of experiments using Ub-Rho110 to validate structural insights. For example, mutations at the DUB active site or in the ubiquitin-binding interface can be rapidly assessed for their functional consequences using Ub-Rho110 fluorescence assays. The substrate has also been used in fluorescence anisotropy and stopped-flow experiments to study binding kinetics and conformational dynamics of DUB catalysis. 4. Assay Design and Optimization 4.1 Buffer and pH Conditions Standard assay buffers for Ub-Rho110 DUB assays typically consist of 50 mM HEPES (pH 7.5), 0.5 mM EDTA, and 1–5 mM DTT or 1 mM TCEP. Some DUBs also require additional components such as NaCl (50–150 mM) for stability or zinc ions (for metalloprotease DUBs). The pH optimum varies by enzyme family: most cysteine DUBs are active over a broad neutral range (pH 6.5–8.0), while metalloprotease DUBs may have different optima. Detergents (e.g., 0.01% Tween-20 or 0.1% BSA) can reduce enzyme adsorption to microplate surfaces and improve assay reproducibility. 4.2 Enzyme Activation and Pre-incubation Cysteine DUBs require reduction of the active-site cysteine for full activity. It is standard practice to pre-incubate recombinant cysteine DUBs with 1–5 mM DTT or 1 mM TCEP for 10–30 minutes at room temperature or 4°C before adding substrate. Some DUBs also require pre-incubation with cofactors (e.g., JAMM DUBs require zinc), and enzyme stabilizers such as BSA or carrier proteins may be needed for enzymes prone to aggregation at low concentrations. 4.3 Substrate Concentration and Linearity To measure initial velocity accurately, it is essential to work in the linear phase of the progress curve, where less than 10–20% of the substrate is consumed. For kinetic parameter determination, substrate concentrations should span at least 0.1–10× KM. Typical working concentrations of Ub-Rho110 range from 0.1 to 10 μM, though some low-KM enzymes may require even lower concentrations to avoid substrate saturation. Inner filter effects, which cause apparent decreases in fluorescence at high substrate concentrations, should also be considered and appropriate correction applied if needed. 4.4 Controls and Quality Metrics Every Ub-Rho110 assay should include the following controls to ensure data quality: • Positive control: Active enzyme at a known, saturating concentration — confirms assay functionality and provides maximum signal. • Negative control (no enzyme): Establishes baseline fluorescence and background hydrolysis rate. • Inhibitor control: A well-characterized DUB inhibitor (e.g., N-ethylmaleimide for cysteine DUBs) should be included to validate inhibitor detection in HTS. • Free Rhodamine 110 standard curve: Used to convert raw fluorescence units to molar product concentrations, enabling calculation of absolute rates. 5. Comparison to Related Substrates 5.1 Ub-AMC (Ubiquitin-7-amino-4-methylcoumarin) Ub-AMC is historically the most widely used DUB fluorogenic substrate. AMC has an excitation/emission of approximately 350/440 nm, placing it in the UV-visible range where interference from compound autofluorescence is more common, particularly in HTS campaigns involving compound libraries. Ub-Rho110 offers several advantages over Ub-AMC: its excitation and emission are in the visible range (496/520 nm), reducing compound interference; Rhodamine 110 has a significantly higher quantum yield; and the signal-to-background ratio is generally superior. However, Ub-AMC remains valuable for mechanistic studies and is still widely available and cost-effective. 5.2 Ub-TAMRA and Other Rhodamine Conjugates Several other rhodamine-based ubiquitin substrates have been developed, including TAMRA (tetramethylrhodamine) conjugates. These alternatives offer distinct spectral properties useful for multiplexing. However, Rhodamine 110 remains the preferred fluorophore for most DUB assay applications due to its superior quantum yield, photostability, and the well-established fluorescence quenching mechanism when conjugated to the ubiquitin C-terminus. 5.3 Diubiquitin Substrates Diubiquitin (Ub-Ub) substrates linked through specific lysine residues (K6, K11, K29, K33, K48, K63) or linear (M1) linkages have been developed to study linkage-selective DUBs. While more informative about linkage preference, they are generally more expensive, more complex to produce, and require gel-based or mass spectrometry-based readouts. Ub-Rho110 remains the substrate of choice for initial activity screening and HTS due to its simplicity, sensitivity, and cost-effectiveness. Substrate: Ub-Rho110 Excitation/Emission: 496/520 nm Signal Enhancement: 100– >1000× HTS Compatibility: Excellent Linkage Selectivity: None (C-terminal) Substrate: Ub-AMC Excitation/Emission: 350/440 nm Signal Enhancement: ~100× HTS Compatibility: Good (UV interference) Linkage Selectivity: None (C-terminal) Substrate: Ub-TAMRA Excitation/Emission: 541/568 nm Signal Enhancement: ~50–80× HTS Compatibility: Good Linkage Selectivity: None (C-terminal) Substrate: Diubiquitin Excitation/Emission: N/A (gel/MS) Signal Enhancement: N/A HTS Compatibility: Poor Linkage Selectivity: Linkage-specific 6. Limitations and Potential Pitfalls 6.1 Pan-DUB Substrate Because Ub-Rho110 reports on total DUB activity rather than the activity of a specific enzyme, it is not inherently selective. In complex biological samples such as cell lysates, the observed signal represents the cumulative activity of all active DUBs present. This limits its use for studying individual DUBs without prior enzyme purification or immunodepletion. 6.2 Compound Interference in HTS Fluorescent compounds in screening libraries can interfere with assay readout through inner filter effects, compound fluorescence, or fluorescence quenching. Counter-screens using free Rhodamine 110 should be performed to identify and triage fluorescent compounds. Absorbance-based compound interference can be addressed by assay miniaturization, shorter incubation times, and the use of fluorescence lifetime or time-resolved fluorescence (TR-FRET) adaptations where available. 6.3 Non-DUB Proteases Although the ubiquitin C-terminal amide bond is highly specific to DUBs under physiological conditions, non-specific proteases present in impure enzyme preparations or crude lysates can cleave the substrate and generate false-positive signals. Protease inhibitor cocktails (excluding DUB inhibitors) can be added to lysate-based assays to minimize this artifact. 6.4 Limited Structural Information The Rhodamine 110 moiety occupies the S1′ exit channel of the DUB active site and may not fully recapitulate interactions seen with natural ubiquitinated substrates. Kinetic parameters measured using Ub-Rho110 may therefore differ from those obtained with more physiologically relevant substrates such as Ub-protein conjugates or diubiquitin chains. Caution should be exercised when extrapolating mechanistic conclusions from Ub-Rho110 data alone. 7. Therapeutic Relevance and Drug Discovery Applications The ubiquitin-proteasome system (UPS) and the DUB enzymes that regulate it are increasingly recognized as important therapeutic targets. Dysregulation of DUB activity has been linked to multiple pathologies: • Cancer: USP7, USP2, USP4, USP14, and UCHL1 are among the DUBs overexpressed or mutated in various cancers, promoting tumor growth by stabilizing oncoproteins or destabilizing tumor suppressors. • Neurodegeneration: UCH-L1 mutations are associated with Parkinson’s disease; USP14 and other DUBs regulate tau and alpha-synuclein turnover. • Viral infections: Several viruses encode their own DUBs or hijack cellular DUBs to evade immune responses, making viral DUBs potential antiviral drug targets. • Inflammatory diseases: OTULIN, CYLD, and A20 are DUBs that negatively regulate NF-κB signaling; their dysregulation contributes to autoimmune and inflammatory conditions. Ub-Rho110 has been central to the discovery and characterization of several DUB inhibitor classes, including covalent inhibitors targeting the active-site cysteine (e.g., vinyl sulfones, propargylamines), non-covalent inhibitors, and PROTAC-based degraders of DUBs. The substrate continues to serve as the primary biochemical validation tool in DUB-targeted drug programs. 8. Emerging Developments and Future Directions 8.1 NEDD8-Rho110 and UBL Substrates Analogous fluorogenic substrates based on ubiquitin-like proteins (UBLs) such as NEDD8-Rho110, SUMO-Rho110, and ISG15-Rho110 have been developed to study the corresponding UBL-processing enzymes (deneddylases, SENPs, and USP18). These substrates enable parallel profiling of multiple UBL pathways using the same assay platform and fluorometer. 8.2 Linkage-Specific Fluorogenic Substrates Efforts to develop linkage-selective fluorogenic substrates have led to diubiquitin-rhodamine conjugates that incorporate Rhodamine 110 at the distal ubiquitin C-terminus within defined chain topologies. These next-generation substrates enable kinetic characterization of linkage-specific DUBs in a fluorescence-based format, combining the selectivity of diubiquitin substrates with the sensitivity of the Rho110 readout. 8.3 Cell-Permeable and In-Cell Applications Although Ub-Rho110 is primarily used in biochemical (cell-free) assays, modified versions with cell-penetrating peptide tags or other cell-delivery strategies are being explored to enable direct measurement of DUB activity in living cells. Such tools would represent a significant advance for pharmacodynamic monitoring of DUB inhibitors in cellular and in vivo models. 8.4 Multiplexed and Microfluidic Assay Formats Integration of Ub-Rho110 assays into microfluidic platforms and droplet-based screening technologies is expanding throughput capabilities and reducing reagent costs. Multiplexed assays combining Ub-Rho110 with orthogonal substrates (using spectrally distinct fluorophores) allow simultaneous profiling of multiple DUBs or UBL pathways in a single well. 9. Summary Ubiquitin-Rhodamine 110 is a versatile, sensitive, and well-validated fluorogenic substrate for the study of DUBs and ubiquitin C-terminal hydrolases. Its combination of high signal enhancement, visible-range fluorescence, compatibility with standard instrumentation, and applicability across a wide range of DUB families and experimental contexts has established it as a foundational reagent in ubiquitin biology research. Below is a consolidated summary of its key attributes: Substrate class: Fluorogenic, C-terminal amide conjugate Fluorophore: Rhodamine 110 (Ex/Em: 496/520 nm) Signal mechanism: Fluorescence dequenching upon DUB-mediated hydrolysis Target enzymes: DUBs (all families), UCHs, UBL-processing enzymes Assay formats: Continuous kinetic, endpoint, HTS (96/384/1536-well) Primary applications: Enzyme activity, HTS, kinetic characterization, lysate profiling Key advantages : High sensitivity, visible-range detection, broad DUB compatibility Key limitations: Pan-DUB substrate, potential compound interference, no linkage selectivity Storage: -80°C, protect from light, single-use aliquots recommended References 1. Dang LC, Melandri FD, Stein RL. Kinetic and mechanistic studies on the hydrolysis of ubiquitin C-terminal 7-amido-4-methylcoumarin by deubiquitinating enzymes. Biochemistry. 1998;37(7):1868–1879. 2. Borodovsky A, Ovaa H, Kolli N, et al. Chemistry-based functional proteomics reveals novel members of the deubiquitinating enzyme family. Chem Biol. 2002;9(10):1149–1159. 3. Komander D, Clague MJ, Urbé S. Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol. 2009;10(8):550–563. 4. Ritorto MS, Ewan R, Perez-Oliva AB, et al. Screening of DUB activity and specificity by MALDI-TOF mass spectrometry. Nat Commun. 2014;5:4763. 5. Mevissen TET, Komander D. Mechanisms of deubiquitinase specificity and regulation. Annu Rev Biochem. 2017;86:159–192. 6. Harrigan JA, Jacq X, Martin NM, Jackson SP. Deubiquitylating enzymes and drug discovery: emerging opportunities. Nat Rev Drug Discov. 2018;17(1):57–78. 7. Clague MJ, Urbé S, Komander D. Breaking the chains: deubiquitylating enzyme specificity begets function. Nat Rev Mol Cell Biol. 2019;20(6):338–352. 8. Sommer S, Weikart ND, Linne U, Mootz HD. Covalent inhibition of SUMO and ubiquitin-specific cysteine proteases by an in situ thiol-alkyne addition. Bioorg Med Chem. 2013;21(9):2422–2431. |
|
Ubiquitin-Rhodamine 110 (Ub-Rho110)
For Research & Development use only. Not for testing and/or use on humans.