| Molecular Formula: | C379H629N105O116S |
| Molecular Weight: | 8544.75 |
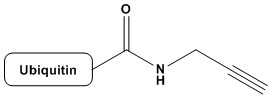
| Ubiquitin-propargylamide (Ub-PA, also referred to as Ub-Prg) is an electrophilic, activity-based probe (ABP) derived from the 76-amino acid protein ubiquitin, in which the C-terminal glycine residue is modified with a propargylamide warhead. This warhead — a terminal alkyne tethered via an amide bond — reacts with the catalytic cysteine residues of deubiquitylating enzymes (DUBs) through a Michael-type addition mechanism, forming a covalent, irreversible adduct. Since its original development in the early 2000s by the Ploegh laboratory, Ub-PA has become one of the most widely deployed tools in ubiquitin biology, enabling the profiling, identification, and functional characterization of active DUBs in cellular lysates, living cells, and whole organisms. 1. Introduction to the Ubiquitin System 1.1 Ubiquitin: Structure and Function Ubiquitin (Ub) is a highly conserved, 76-amino acid protein (molecular weight ~8.6 kDa) found in virtually all eukaryotic organisms. Its sequence is remarkably conserved across evolution — yeast and human ubiquitin differ in only 3 of 76 residues — reflecting its fundamental biological importance. The three-dimensional structure of ubiquitin adopts a compact beta-grasp fold (also called ubiquitin-like or UBL fold), characterized by a five-stranded antiparallel beta-sheet, a short 3.5-turn alpha-helix, and a C-terminal tail ending in the di-glycine motif Gly75-Gly76. This flexible C-terminal tail is central to ubiquitin’s function: the terminal carboxylate of Gly76 is conjugated to substrate proteins through an isopeptide bond with lysine residues, or in some cases through thioester intermediates or N-terminal amine bonds. Ubiquitination — the covalent attachment of ubiquitin to protein substrates — is a central post-translational modification (PTM) orchestrating diverse cellular processes. These include proteasomal and lysosomal protein degradation, DNA damage response and repair, cell cycle progression, immune signaling, endosomal sorting, autophagy, and the regulation of kinase activity. The functional consequences of ubiquitination are largely dictated by the topological architecture of the ubiquitin signal: monoubiquitination, multi-monoubiquitination, and the formation of polyubiquitin chains linked through any of ubiquitin’s seven internal lysine residues (K6, K11, K27, K29, K33, K48, K63) or through the N-terminal methionine (M1, linear chains). K48-linked polyubiquitin chains are the canonical signal for proteasomal degradation, while K63-linked chains are associated with DNA repair, endosomal trafficking, and NF-κB signaling. M1/linear chains play specialized roles in inflammatory signaling through the LUBAC complex. K11-linked chains serve cell cycle functions and are part of the APC/C-mediated degradation pathway. K27, K29, and K33 linkages remain less well characterized, while K6 chains have been implicated in mitophagy and genome integrity. The remarkable signaling diversity of ubiquitin arises in part from the distinct structural conformations adopted by different chain types, and from the specialized reader proteins (UBDs, ubiquitin-binding domains) that decode these signals. 1.2 The Ubiquitin Conjugation Cascade Ubiquitination is catalyzed by a hierarchical enzymatic cascade comprising three classes of enzymes. E1 ubiquitin-activating enzymes initiate the process by adenylating the C-terminal glycine of ubiquitin and forming a high-energy thioester bond between their catalytic cysteine and ubiquitin. In humans, two E1 enzymes (UBA1 and UBA6) are responsible for this activation step. E2 ubiquitin-conjugating enzymes (of which there are ~40 in humans) then accept the activated ubiquitin from E1 via transthioesterification. Finally, E3 ubiquitin ligases (with over 600 predicted in humans) mediate the transfer of ubiquitin to specific substrate proteins, either by direct catalysis (HECT and RBR E3s) or by positioning the E2-Ub intermediate near the substrate (RING E3s). This enzymatic cascade is tightly regulated and highly substrate-specific, allowing ubiquitin to function as a dynamic and reversible signaling molecule. The reversibility of ubiquitination is provided by deubiquitylating enzymes (DUBs), which constitute the opposing arm of the ubiquitin system and are the primary biochemical targets of Ub-PA. 2. Deubiquitylating Enzymes (DUBs): Classification and Catalytic Mechanisms 2.1 Overview and Biological Roles Deubiquitylating enzymes (DUBs, also called deubiquitinases) are proteases that catalyze the hydrolysis of isopeptide bonds between ubiquitin and substrate proteins, or within polyubiquitin chains. The human genome encodes approximately 100 DUBs, which perform a range of critical cellular functions: recycling ubiquitin from proteasome-targeted substrates, editing polyubiquitin chain topology, rescuing proteins from degradation, processing ubiquitin precursors (linear ubiquitin gene products), and regulating receptor trafficking, gene expression, and DNA repair pathways. DUBs are classified into seven major families based on their catalytic domain architecture. Six of these — the ubiquitin-specific proteases (USPs), the ubiquitin C-terminal hydrolases (UCHs), the ovarian tumor proteases (OTUs), the Machado-Joseph disease protein domain proteases (MJDs/Josephins), the MINDY (motif interacting with Ub-containing novel DUB) family, and the ZUP1/ZUFSP family — are cysteine proteases. The seventh family, the JAMM/MPN metalloprotease domain-containing enzymes, uses a zinc-coordinated water molecule to catalyze hydrolysis, analogous to classical metalloproteases. 2.2 Catalytic Mechanisms Cysteine-dependent DUBs employ a catalytic triad (Cys-His-Asp/Asn, analogous to papain-family proteases) to cleave isopeptide bonds. The catalytic cysteine acts as a nucleophile, attacking the isopeptide carbonyl carbon to form an acyl-enzyme intermediate, with concomitant departure of the amine of the substrate lysine residue. The intermediate is subsequently hydrolyzed by a water molecule, releasing ubiquitin with a free C-terminal carboxylate and regenerating the active enzyme. A critical feature of most cysteine DUBs in their unliganded state is that the catalytic cysteine exists in a partially oxidized or otherwise inactive conformation. Structural studies have revealed that the active site undergoes conformational rearrangements upon ubiquitin binding — a phenomenon sometimes called ‘ubiquitin-induced activation.’ This structural plasticity has important implications for the selectivity of activity-based probes like Ub-PA, since only properly folded, active-state DUBs will react with the probe. JAMM metalloprotease DUBs, including AMSH, AMSH-LP, BRCC36 (in the BRISC and BRCA1-BRCA2-containing complexes), and CSN5 (in the COP9 signalosome), utilize a different catalytic mechanism. A zinc ion coordinated by conserved His-x-His-x(n)-Asp residues activates a water molecule for nucleophilic attack on the isopeptide bond. Because JAMM enzymes lack a catalytic cysteine, cysteine-reactive probes like Ub-PA do not label JAMM DUBs. This represents an important selectivity characteristic of the probe. 3. Activity-Based Protein Profiling (ABPP): Conceptual Framework Activity-based protein profiling (ABPP) is a chemical biology strategy that uses small molecule or protein-based probes to covalently and irreversibly label the active forms of specific enzyme families in complex biological samples. Unlike antibody-based or transcriptomic approaches, ABPP directly reports on the catalytically active fraction of an enzyme population, providing information inaccessible through conventional proteomics. An activity-based probe (ABP) consists of three functional elements. First, a recognition element (or binding unit) that confers selectivity for a specific enzyme class — in the case of Ub-PA, the ubiquitin protein scaffold directs the probe toward DUBs and other ubiquitin-binding enzymes. Second, a reactive group (or warhead) that covalently modifies the enzyme’s active site upon binding — in Ub-PA, this is the propargylamide moiety. Third, a reporter tag for downstream detection and identification — which in Ub-PA can be incorporated via chemical conjugation or recombinant expression strategies. ABPP has been transformative for the characterization of serine hydrolases, cysteine proteases, kinases, and numerous other enzyme families. The development of Ub-PA adapted this powerful approach to the ubiquitin system, enabling the global profiling of DUB activity states in complex proteomes. 4. Design and Synthesis of Ubiquitin-Propargylamide 4.1 Rational Design Principles The design of Ub-PA was predicated on several key principles derived from knowledge of DUB biochemistry and mechanism. First, since DUBs recognize and bind ubiquitin prior to catalysis, a probe based on the full-length ubiquitin scaffold would be expected to bind the DUB active site with high affinity and selectivity, mimicking the natural substrate. Second, a warhead positioned at the C-terminus of ubiquitin — the site that occupies the active site of DUBs — would be ideally placed to react with the catalytic residue. Third, the warhead must be reactive toward the catalytic nucleophile (cysteine thiol for most DUBs) without being excessively reactive toward off-target nucleophiles, ensuring acceptable selectivity in complex biological matrices. Propargylamide fulfills these criteria elegantly. The propargylamide group (HC≡C-CH2-NH-CO-) is a Michael acceptor: the alkyne, activated by the adjacent amide carbonyl through conjugation, can undergo addition of nucleophilic thiols via a thiol-yne reaction. The reactivity is modulated by the amide — sufficiently electrophilic to react with the highly nucleophilic active-site cysteine in the DUB catalytic pocket, yet not so reactive as to alkylate non-enzymatic thiols indiscriminately. The reaction produces a vinyl thioether linkage, forming a stable, essentially irreversible adduct. 4.2 Synthesis Strategies The synthesis of Ub-PA has evolved considerably since its initial development. The original approach, pioneered by Borodovsky et al. (2002) and Hemelaar et al. (2004), relied on expressed protein ligation (EPL) — a semisynthetic strategy that combines recombinant protein expression with synthetic peptide chemistry. In the EPL approach, a ubiquitin(1-75)-intein fusion protein is expressed recombinantly in E. coli. The C-terminal intein undergoes N-to-S acyl shift under mildly reductive conditions, generating a ubiquitin(1-75)-thioester intermediate. This thioester is then reacted with a synthetic propargylamine-containing dipeptide (H-Gly-propargylamide) under native chemical ligation conditions, installing the propargylamide warhead at the C-terminus. The resulting full-length ubiquitin-propargylamide is purified by HPLC, with typical yields of 1-10 mg per liter of bacterial culture depending on expression conditions. An alternative and increasingly popular approach uses sortase-mediated transpeptidation. Sortase A from Staphylococcus aureus recognizes the LPXTG motif and mediates transpeptidation between the threonine carbonyl and an N-terminal glycine-containing nucleophile. By engineering a sortase recognition sequence at the C-terminus of ubiquitin, the propargylamide group can be appended through a sortase-mediated ligation step, avoiding the need for intein technology. Fully synthetic approaches using solid-phase peptide synthesis (SPPS) have also been reported for the synthesis of ubiquitin analogs, including Ub-PA, though the large size of ubiquitin (76 residues) makes this approach technically demanding. More recently, amber suppression/unnatural amino acid incorporation strategies have been explored to introduce the propargylamide at specific positions with high site-specificity. 4.3 Incorporation of Reporter Tags To facilitate detection and identification of labeled DUBs, Ub-PA is commonly equipped with reporter tags. These include fluorophores (such as TAMRA, Cy3, or BODIPY) for in-gel fluorescence scanning, biotin for streptavidin-mediated enrichment and mass spectrometric identification, and FLAG or HA epitope tags for immunoblot detection. In the original EPL synthesis, biotin is conveniently incorporated into the C-terminal propargylamine-containing peptide, yielding Ub-Prg-biotin as a single synthetic step. Alternatively, copper-catalyzed azide-alkyne cycloaddition (CuAAC/’click chemistry’) can be used to install reporter groups post-labeling: the alkyne of the propargylamide, after reaction with the DUB cysteine, leaves the unreacted alkyne on the distal side accessible for click chemistry — though the chemistry of this has nuances depending on the adduct structure. 5. Mechanism of Covalent Inhibition 5.1 Molecular Mechanism of Reaction The reaction between Ub-PA and a cysteine DUB proceeds through a well-defined mechanism. The catalytic cysteine thiol (in its reactive thiolate form, pKa-depressed by the adjacent histidine) is positioned at the bottom of the active site cleft, which is shaped to recognize the C-terminal tail of ubiquitin. Upon binding of Ub-PA to the DUB active site, the propargylamide warhead is positioned immediately adjacent to the catalytic cysteine. The thiolate performs a nucleophilic 1,4-addition (Michael addition) to the alpha,beta-unsaturated system of the propargylamide. Specifically, the sulfur attacks the beta-carbon of the propargyl group (the internal alkyne carbon), generating a vinyl sulfide intermediate that tautomerizes to a stable vinyl thioether. This vinyl thioether is not susceptible to hydrolysis under physiological conditions, rendering the DUB-Ub-PA complex essentially irreversible. The formation of the covalent adduct results in a mass increase of approximately 8,631 Da (the mass of Ub-PA minus the proton donated to the leaving group), which is readily detectable by intact protein mass spectrometry. An important mechanistic feature is the selectivity of the propargylamide for the active-site cysteine over other cellular thiols. The active-site cysteine of DUBs is uniquely reactive due to its low pKa (stabilized by the His-Asp catalytic triad), conferring enhanced nucleophilicity compared to ordinary cysteine residues. Furthermore, the proximal positioning of the warhead within the active site pocket — enforced by the non-covalent ubiquitin-DUB interaction — provides a proximity effect that dramatically accelerates the intra-complex reaction relative to bimolecular reactions with non-active-site thiols. 5.2 Selectivity for Active Versus Inactive DUBs One of the most valuable properties of Ub-PA is its selectivity for active, properly folded DUBs. Enzymes whose active site cysteine has been inactivated — through oxidation to sulfenic acid, irreversible overoxidation to sulfinic or sulfonic acid, glutathionylation, S-nitrosation, or through active site mutation — do not react with Ub-PA. This property makes Ub-PA a true activity reporter, rather than merely an abundance reporter. By comparing Ub-PA labeling profiles between cell states (e.g., before and after drug treatment, between healthy and disease states), changes in DUB activity can be identified that would be invisible to transcriptomic or total protein abundance measurements. Similarly, DUBs that are conditionally active only within multiprotein complexes (e.g., BRCC36 in BRISC/BRCA1 complexes, or MINDY family members) may show differential labeling in intact cell lysates versus purified fractions, providing information about the regulation of DUB activity by protein-protein interactions. 6. Structural Biology of Ub-PA–DUB Complexes X-ray crystallography and cryo-electron microscopy of DUBs in covalent complex with Ub-PA have provided extraordinary structural insights into DUB catalysis, substrate recognition, and the molecular basis of activity-based probe selectivity. These structures also form the foundation for structure-based drug design efforts targeting DUBs. 6.1 Key Crystal Structures The first reported structure of a DUB–Ub-PA complex was the UCH-L3–Ub-PA complex (Misaghi et al., 2005), which confirmed the active-site mechanism and revealed how the C-terminal tail of ubiquitin threads through a crossover loop characteristic of the UCH family. Subsequent structures have illuminated diverse DUBs from all cysteine-dependent families. Landmark structures include the USP7 (HAUSP)–Ub-PA complex, which revealed the ‘palm-fingers-thumb’ architecture of USP family DUBs and how the catalytic triad is reoriented upon ubiquitin binding — illustrating the concept of substrate-induced active site assembly. The USP2–Ub-PA and USP21–Ub-PA structures confirmed that USP family members share this general mechanism despite highly divergent primary sequences outside the catalytic domain. OTU family structures in complex with Ub-PA, including OTUB1–Ub-PA, OTUD3–Ub-PA, and vOTU (from Crimean-Congo hemorrhagic fever virus)–Ub-PA, have been particularly informative. These structures revealed a strikingly distinct active site architecture from USPs, including the OTU-specific catalytic cysteine, histidine, and variable acidic residue arrangement. The viral OTU structures have illuminated the evolutionary hijacking of the ubiquitin system by pathogenic viruses and have informed the design of antiviral therapeutic strategies. MINDY family DUBs, only recently discovered, have also been crystallized in complex with Ub-PA, revealing a novel catalytic fold with a non-canonical cysteine-histidine catalytic dyad and selectivity for K48-linked chains. The ZUP1/ZUFSP family, characterized by selectivity for K63-linked chains, similarly has been structurally characterized using Ub-PA-based tools. 6.2 Insights into Selectivity and Allostery An important insight from structural studies of DUB–Ub-PA complexes is that DUBs recognize multiple surfaces of ubiquitin beyond just the C-terminal tail. The hydrophobic patch centered on Ile44 of ubiquitin is a critical recognition epitope for many DUBs (as well as UBDs in general). Mutations at this surface (e.g., Ile44Ala) dramatically reduce DUB activity and Ub-PA reactivity, providing a biochemical tool to probe the importance of this interface. Many structures have also revealed allosteric mechanisms: conformational changes propagating from the ubiquitin-binding interface to the active site, ‘switching on’ the catalytic cysteine nucleophilicity. This structural plasticity — where DUBs exist in low-activity apo states and high-activity holo/substrate-bound states — is a key selectivity determinant for Ub-PA, as only enzymes capable of adopting the active conformation will react with the probe. 7. Applications in DUB Profiling and Target Identification 7.1 Proteome-Wide DUB Profiling The seminal application of Ub-PA, reported by Borodovsky et al. (2002), established the concept of DUB profiling: incubation of Ub-PA with cell lysates, followed by SDS-PAGE and visualization of radiolabeled or biotin-tagged products, reveals a characteristic pattern of labeled DUBs that serves as a proteome-wide snapshot of DUB activity. This approach immediately identified several novel DUBs that were not previously recognized as such. Combined with mass spectrometry — either by in-gel digestion of Ub-PA-reactive bands or by streptavidin pull-down of biotin-tagged Ub-PA labeled proteins followed by LC-MS/MS — this approach has enabled the unbiased identification of dozens of DUBs in diverse cell types and conditions. The technique is particularly powerful when combined with quantitative proteomics approaches such as SILAC (stable isotope labeling by amino acids in cell culture) or TMT (tandem mass tag) multiplexing, allowing direct quantitative comparison of DUB activities across multiple conditions simultaneously. 7.2 Identification of Novel DUBs Ub-PA-based profiling has contributed directly to the discovery and validation of novel DUBs. Prior to the advent of ABPP approaches, DUB identification relied primarily on bioinformatics (sequence homology to known catalytic domains) and conventional biochemical assays with model substrates. Ub-PA profiling expanded this by providing direct biochemical evidence of catalytic activity for previously uncharacterized proteins. Notable examples include the initial characterization of several MINDY (MINDY1-4) family members, whose DUB activity was confirmed through Ub-PA reactivity before their crystal structures were determined. Similarly, the ZUFSP/ZUP1 enzyme, discovered as a K63-selective DUB, was confirmed as a bona fide DUB in part through its reactivity with Ub-PA. 7.3 Studying DUB Regulation Beyond identification, Ub-PA profiling reveals how DUB activities are regulated. Differential profiling — comparing Ub-PA labeling between cell lines, genetic knockouts, or drug-treated samples — can reveal DUBs whose activity is specifically regulated under particular conditions. For example, DUBs that require specific protein partners for full activity may show enhanced or reduced labeling in cells lacking those partners. Ub-PA has been used to study the activation of DUBs upon proteasome inhibition: treatment of cells with proteasome inhibitors (e.g., MG132, bortezomib) leads to ubiquitin depletion and compensatory changes in DUB activity profiles that are detectable by Ub-PA labeling. Similarly, changes in DUB activities during mitosis, apoptosis, DNA damage responses, and viral infections have been characterized using this approach. 8. Ub-PA in Cell-Based and In Vivo Studies 8.1 Cell-Permeable Ub-PA Variants A limitation of early Ub-PA studies was that the probe was large (>8.6 kDa) and therefore could not passively diffuse across plasma membranes. Studies were thus primarily performed in cell lysates, limiting the ability to characterize DUB activities in intact cells where compartmentalization and regulatory protein complexes are preserved. To address this, several strategies for cell-permeable delivery of Ub-PA have been developed. One approach uses cell-penetrating peptides (CPPs), such as polyarginine or TAT peptide sequences, conjugated to Ub-PA to facilitate cellular uptake. Microinjection, electroporation, and bead loading are alternative physical approaches that have been used to introduce Ub-PA into living cells. Fusion to endosome-disruptive agents can enhance cytosolic delivery following endocytic uptake. A conceptually distinct approach uses smaller ubiquitin-based probes — such as C-terminal electrophilic warhead-modified ubiquitin fragments — that retain sufficient DUB binding affinity while being more amenable to cellular uptake. However, these smaller constructs generally show reduced selectivity compared to full-length Ub-PA. 8.2 In Vivo Applications and Organismal Studies Ub-PA has been applied in whole-organism contexts using both injection and genetic approaches. In zebrafish and Drosophila models, fluorescently tagged Ub-PA has been used to visualize active DUB populations in developing embryos, revealing spatiotemporal regulation of DUB activity during development. In mouse models, Ub-PA-based tools have been used to characterize DUB activity profiles in specific tissues and in disease models including cancer, neurodegeneration, and infection. The ability to compare DUB activity profiles between normal and diseased tissues has identified DUBs with potential roles in disease pathogenesis. 9. Contributions to Drug Discovery 9.1 DUBs as Therapeutic Targets DUBs represent an increasingly attractive class of drug targets. The ubiquitin-proteasome system (UPS) has already been clinically validated: proteasome inhibitors (bortezomib, carfilzomib, ixazomib) are approved therapies for multiple myeloma. DUBs, however, offer potentially superior specificity over the proteasome itself, as individual DUBs have discrete, non-redundant substrate specificities. Among the most intensively pursued DUB targets are USP7 (HAUSP, implicated in p53 pathway regulation and oncology), USP1 (DNA repair), USP2, USP11, USP14, UCHL5 (proteasome-associated DUBs), and CYLD (tumor suppressor). Several DUB-targeted compounds have entered clinical development. VLS-1488 (a USP1 inhibitor) and other USP1-targeting agents have shown promise in BRCA-mutant cancers by exploiting synthetic lethality. USP7 inhibitors are under active investigation for solid tumors and hematological malignancies. 9.2 Ub-PA in DUB Inhibitor Development Ub-PA plays multiple roles in DUB drug discovery pipelines. First, it is used as a competition probe: a DUB inhibitor that occupies the active site will prevent Ub-PA from reacting, and the degree of competition can be quantified by gel-based or LC-MS-based assays. This is sometimes called a competitive ABPP or chemoproteomics approach. By running such competition assays in cell lysates or living cells (using cell-permeable Ub-PA delivery strategies), researchers can assess inhibitor potency and selectivity across the entire expressed DUB proteome simultaneously. Second, Ub-PA-mediated crystallography provides the structural templates essential for structure-based inhibitor design. Structures of DUBs in complex with Ub-PA reveal the active site geometry, S1/S1′ ubiquitin binding pockets, and peripheral contacts that can be exploited for inhibitor design. Third, in target engagement assays for clinical candidates, Ub-PA labeling (with the probe as a molecular tool) can confirm that a drug candidate is occupying the intended DUB active site in cell-based or patient-derived tissue samples, informing on pharmacodynamic biomarkers. 10. Variants, Analogs, and Extended Probe Families 10.1 Chain Topology-Selective Probes The original Ub-PA probe consists of a single ubiquitin molecule. To probe DUBs with selectivity for specific polyubiquitin chain linkages, diubiquitin-based probes have been developed. Diubiquitin-PA (diUb-PA) probes consist of two ubiquitin molecules linked via a specific isopeptide bond (e.g., K48-linked, K63-linked, M1-linked, or K11-linked), with the propargylamide warhead at the C-terminus of the distal ubiquitin. These diUb-PA probes preferentially label DUBs with selectivity for the corresponding chain topology, enabling the profiling of linkage-selective DUBs such as the K63-selective USP18 and CYLD, the K48-selective MINDY family, and the M1-selective OTULIN. Extension to triubiquitin and longer chain probes has also been reported, with increasingly complex synthesis challenges. 10.2 Ubiquitin-Like Modifier Probes The success of Ub-PA inspired the development of analogous probes based on ubiquitin-like modifiers (UBLs), including NEDD8, SUMO (SUMO-1, SUMO-2/3), ISG15, FAT10, UFM1, and URM1. These UBL-PA probes enable the profiling of the corresponding deconjugating enzymes: NEDDylases (deneddylases such as CSN5/SENP8 and NEDP1), sentrin/SUMO-specific proteases (SENPs), ISG15-specific USP18, FAT10-directed DUBs, and so on. These expanded probe families have collectively enabled a comprehensive view of the UBL-deconjugase landscape and have revealed cross-reactivity between some DUBs and multiple UBL substrates, with important implications for signaling specificity. 10.3 Warhead Variants Beyond propargylamide, numerous alternative C-terminal warheads have been reported for ubiquitin-based activity-based probes. Vinyl sulfones (Ub-VS, the first-generation DUB ABPP tool, introduced even before Ub-PA) react by 1,4-addition with the catalytic cysteine but produce a different adduct structure. Vinyl methyl esters (Ub-VME) similarly react with cysteine DUBs. Dehydroalanine-based probes, propargyl-Gly probes, and various Michael acceptor-modified C-terminal residues have been evaluated. Each warhead has distinct reactivity profiles: Ub-VS tends to be more reactive but less selective, while Ub-PA is generally regarded as the optimal balance of reactivity, selectivity, and cross-reactivity with the broadest range of cysteine DUBs. Photoaffinity labeling approaches using diazirine or benzophenone-modified ubiquitin have also been developed to capture weaker or more transient DUB-ubiquitin interactions. 11. Comparison with Alternative DUB Probes and Assays Ub-PA occupies a distinct and complementary niche among approaches for studying DUB activity. Fluorogenic substrate assays using Ub-AMC (ubiquitin-7-amino-4-methylcoumarin) or Ub-Rho110 (ubiquitin-rhodamine) provide real-time kinetic information about DUB activity but require purified DUBs and do not report directly on the active DUB fraction in complex proteomes. FRET-based diubiquitin substrates allow linkage-selective activity measurement in real time. In contrast to these fluorogenic substrates, Ub-PA covalently captures DUBs, enabling proteome-wide identification by mass spectrometry. However, Ub-PA does not provide kinetic (rate) information — it is a stoichiometric label rather than a turnover-dependent reporter. Gel-based assays also lack the temporal resolution of fluorogenic methods. Non-covalent approaches, such as ubiquitin-derived inhibitors or competitive binding assays using fluorescence polarization, provide complementary information about DUB occupancy without covalent modification. Proximity-based biotinylation (BioID/TurboID) fused to ubiquitin can capture DUBs in native cellular contexts without requiring probe delivery. The integration of Ub-PA-based ABPP with quantitative mass spectrometry currently represents the most informative single approach for comprehensive DUB profiling across complex proteomes, and remains the reference method against which alternatives are benchmarked. 12. Limitations and Caveats Despite its power, Ub-PA has several important limitations that must be considered when interpreting experimental data. Membrane impermeability is a fundamental constraint. As noted, the large size of Ub-PA (>8.6 kDa) prevents passive membrane diffusion, restricting most applications to cell lysates or requiring specialized delivery strategies for intact cell experiments. This means that DUBs whose activity is dependent on membrane context (e.g., those associated with the ER membrane or lysosomal surface) may be underrepresented in lysate-based profiling experiments. JAMM metalloprotease DUBs are completely invisible to Ub-PA, as their catalytic mechanism does not involve a reactive cysteine. Profiling of JAMM DUBs requires complementary approaches, including active-site metalloprotease probes or JAMM-selective inhibitor-competition assays. DUBs requiring specific protein cofactors or post-translational modifications for activity may show altered reactivity in cell lysates compared to their in vivo activity state, since the dilution of lysate disrupts protein complexes and washes away modifying enzymes. Comparison of Ub-PA labeling profiles in intact versus lysed cells (using delivery strategies) is therefore informative. Active site cysteine oxidation, a common event during cell lysis under aerobic conditions, can lead to underestimation of DUB activity. The inclusion of reducing agents (DTT or TCEP) during lysis can partially mitigate this, though some DUBs are irreversibly inactivated by oxidation. This issue affects all cysteine-reactive ABPP probes. The probe does not distinguish between catalytic events — it simply reports on the presence of an active, reactive catalytic cysteine. Some ‘DUBs’ that react with Ub-PA may primarily function as ubiquitin-binding proteins with low catalytic activity, or may have non-DUB physiological roles. Careful biochemical validation of hits identified by Ub-PA profiling is therefore important. 13. Future Directions Several exciting directions are emerging in the Ub-PA and DUB ABPP field. The development of cell-permeable, full-length Ub-PA remains an important goal. Advances in protein delivery technologies — including engineered cell-penetrating protein carriers, lipid nanoparticle delivery, and in situ probe expression via mRNA delivery — may eventually enable routine in-cell Ub-PA profiling. Integration with advanced mass spectrometry workflows, including data-independent acquisition (DIA) proteomics and single-cell proteomics, promises to extend the sensitivity and throughput of Ub-PA-based profiling to previously inaccessible sample types and quantities. Single-cell DUB activity profiling could reveal heterogeneity in DUB activity states within cell populations. Targeted covalent inhibitor design inspired by the Ub-PA warhead chemistry continues to advance. Electrophilic fragments derived from the propargylamide warhead, in combination with ubiquitin-mimicking recognition elements, offer a modular design platform for highly selective DUB covalent inhibitors. The success of covalent drugs in other protease families (e.g., KRAS G12C inhibitors, BTK inhibitors) has renewed interest in targeted covalency as a drug design strategy. The expanding catalog of UBL probes — including those based on FAT10, UFM1, and URM1, for which few deconjugases have been biochemically characterized — will continue to reveal new members of the deconjugase landscape. Probes that can simultaneously profile multiple UBL systems will be powerful tools for understanding UBL cross-talk. Finally, advances in cryo-EM are enabling structural characterization of DUB complexes at resolutions previously accessible only to X-ray crystallography, without the requirement for crystal growth. This will accelerate structural studies of large, multi-subunit DUB complexes (e.g., POH1/Rpn11 in the 26S proteasome lid) that have been refractory to crystallization but are amenable to cryo-EM characterization using Ub-PA-crosslinked samples. 14. Conclusions Ubiquitin-propargylamide (Ub-PA) represents one of the most significant chemical biology tools to emerge from the field of ubiquitin research. By combining the exquisite binding specificity of the ubiquitin protein scaffold with the targeted reactivity of the propargylamide warhead, Ub-PA achieves selective, covalent, and irreversible modification of active cysteine-dependent DUBs across complex biological matrices. Since its introduction, Ub-PA and its derivatives have catalyzed major advances in DUB biology: enabling proteome-wide DUB activity profiling, facilitating the structural characterization of DUB–ubiquitin complexes, providing competitive tools for DUB inhibitor development, and expanding the toolkit for studying the broader UBL landscape. The probe’s impact extends beyond basic research: Ub-PA-based chemoproteomics approaches are now embedded in pharmaceutical DUB drug discovery pipelines, where they serve as indispensable tools for inhibitor selectivity profiling, target engagement assessment, and mechanism-of-action studies. As the therapeutic importance of DUBs continues to grow — reflected in the clinical advancement of DUB-targeted agents — the relevance of Ub-PA as a research and translational tool will only increase. While limitations exist — notably the impermeability to intact membranes, the inability to profile JAMM metalloproteases, and the requirement for specialized delivery approaches for cell-based studies — ongoing technological advances in probe delivery, mass spectrometry, and structural biology continue to expand the reach of Ub-PA-based approaches. Taken together, Ub-PA stands as an enduring exemplar of rational chemical probe design and a cornerstone of the chemical biology of ubiquitin. References 1. Borodovsky A, Ovaa H, Kolli N, et al. (2002). Chemistry-based functional proteomics reveals novel members of the deubiquitinating enzyme family. Chemistry & Biology, 9(10), 1149–1159. [Landmark: first description of Ub-PA and DUB profiling by ABPP] 2. Hemelaar J, Borodovsky A, Bhatt BM, et al. (2004). Specific and covalent targeting of conjugating and deconjugating enzymes of ubiquitin-like proteins. Molecular and Cellular Biology, 24(1), 84–95. [Extended Ub-PA synthesis and validation] 3. Misaghi S, Galardy PJ, Meester WJ, et al. (2005). Structure of the ubiquitin hydrolase UCH-L3 complexed with a suicide substrate. Journal of Biological Chemistry, 280(2), 1512–1520. [First DUB–Ub-PA crystal structure] 4. Komander D, Clague MJ, Urbé S. (2009). Breaking the chains: structure and function of the deubiquitinases. Nature Reviews Molecular Cell Biology, 10(8), 550–563. [Comprehensive DUB review] 5. Ekkebus R, van Kasteren SI, Kulathu Y, et al. (2013). On terminal alkynes that can react with active-site cysteine nucleophiles in proteases. Journal of the American Chemical Society, 135(8), 2867–2870. [Mechanistic characterization of propargylamide warhead] 6. de Jong A, Witting K, Ekkebus R, et al. (2012). Analysis of Ser, Cys and Thr ubiquitin and NEDD8 C-terminal electrophilic probes. Bioorganic & Medicinal Chemistry, 20(17), 5229–5236. 7. Mulder MP, Witting K, Berlin I, et al. (2016). A cascading activity-based probe sequentially targets E1–E2–E3 ubiquitin enzymes. Nature Chemical Biology, 12(7), 523–530. 8. Hewings DS, Flygare JA, Bogyo M, Bhatt AP. (2017). Activity-based probes for the ubiquitin conjugation–deconjugation machinery: new chemistries, new tools, and new insights. FEBS Journal, 284(10), 1557–1580. 9. Hermanns T, Woiwode I, Guerriero G, et al. (2018). An evolutionary approach to systematic discovery of novel deubiquitinases targeting diverse ubiquitin and ubiquitin-like signals. Proceedings of the National Academy of Sciences, 115(21), E4798–E4807. 10. Ovaa H, Ploegh HL. (2004). Probing the ubiquitin-proteasome system with activity-based probes. Trends in Biochemical Sciences, 29(11), 596–601. 11. Mevissen TET, Hospenthal MK, Geurink PP, et al. (2013). OTU deubiquitinases reveal mechanisms and specificity in ubiquitin chain recognition. Cell, 154(1), 169–184. 12. Abdul Rehman SA, Kristariyanto YA, Choi SY, et al. (2016). MINDY-1 is a member of an evolutionarily conserved and structurally distinct new family of deubiquitinating enzymes. Molecular Cell, 63(1), 146–155. [MINDY family discovery, Ub-PA characterization] 13. Haahr P, Borgermann N, Guo X, et al. (2018). ZUFSP deubiquitylates K63-linked polyubiquitin chains to promote genome stability. Molecular Cell, 70(1), 165–174. [ZUP1/ZUFSP discovery] 14. Geurink PP, El Oualid F, Jonker A, et al. (2012). A small-molecule activity-based probe for the ubiquitin isopeptidase UCH-L1. ChemBioChem, 13(2), 293–297. 15. Kategaya L, Di Lello P, Rougé L, et al. (2017). USP7 small-molecule inhibitors interfere with ubiquitin binding. Nature, 550(7677), 534–538. |
|
Ubiquitin-Propargylamide (Ub-PA or Ub-Prg)
For Research & Development use only. Not for testing and/or use on humans.