| Synonym: | L-Photo-Methionine |
| CAS #: | 851960-68-4 |
| Molecular Formula: | C6H11N3O2 |
| Molecular Weight: | 157.2 |
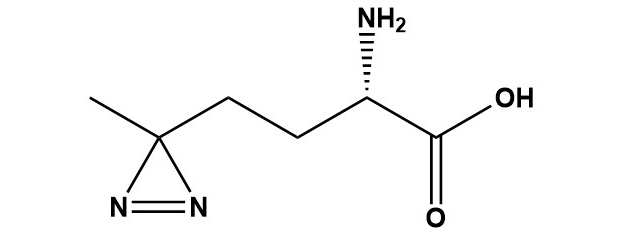
| L-Photo-Methionine (CAS #: 851960-68-4), also commonly referred to as L-Photo-Met, is a synthetic non-canonical amino acid analogue of L-methionine bearing a diazirine ring in place of the natural thioether side chain. First described in 2005 by Suchanek, Radzikowska, and Thiele at the Max Planck Institute of Molecular Cell Biology and Genetics, L-Photo-Met represented a paradigm shift in the study of protein-protein interactions (PPIs) by enabling metabolic incorporation of a photo-crosslinking moiety directly into newly synthesized cellular proteins without requiring genetic code expansion or bulky chemical modifications. Upon irradiation with near-UV light (~355–365 nm), the diazirine ring undergoes photolysis to generate a highly reactive carbene intermediate that inserts into adjacent C-H, N-H, and O-H bonds, forming covalent adducts with nearby protein residues or interaction partners. This approach allows the capture of transient, low-affinity, and otherwise difficult-to-detect protein interactions in their native cellular context. 1. Introduction Protein-protein interactions (PPIs) form the operational backbone of virtually all cellular processes. From signal transduction and gene regulation to vesicle trafficking and immune recognition, the precise, dynamic, and often transient nature of these interactions underpins cell biology. Despite their fundamental importance, studying PPIs in intact cells has remained a major technical challenge. Classical techniques such as co-immunoprecipitation (Co-IP), yeast two-hybrid assays, and affinity chromatography are powerful but suffer from well-known limitations: they frequently cannot capture transient or low-stoichiometry interactions, require disruption of the cellular environment, and may lose weak-binding partners during wash steps. Photoaffinity labeling (PAL) provides an elegant solution to many of these problems by allowing researchers to covalently ‘freeze’ non-covalent, transient interactions using light. Upon UV irradiation, a photoreactive group on or near the probe molecule generates a reactive intermediate that forms a covalent bond with adjacent molecules, permanently recording a molecular interaction that may last only microseconds in the native state. Since the pioneering work of Westheimer in 1962, multiple classes of photoreactive groups have been developed, including aryl azides, benzophenones, and diazirines. Among these, diazirine-based reagents have emerged as the preferred choice for many applications due to their small size, wavelength-dependent selectivity (activatable at 355–365 nm with minimal cell damage), and the high reactivity of the carbene intermediate they produce. L-Photo-Methionine sits at the intersection of these advantages and a key biological insight: cells can be tricked into metabolically incorporating this amino acid analogue into nascent proteins, distributing the photocrosslinker throughout the proteome in a way that is impossible to achieve with conventional exogenous probes. 2. Chemical Identity and Physical Properties 2.1 Structural Overview L-Photo-Methionine is a synthetic amino acid in which the sulfur-containing side chain of natural L-methionine (-CH2-CH2-S-CH3) has been replaced by a diazirine-bearing substituent (-CH2-CH2-C(=N2)-CH3). The diazirine group consists of a three-membered ring containing one carbon and two nitrogen atoms, with the two nitrogens connected by a double bond. This compact, strained ring is the photoreactive element responsible for the compound’s unique properties. Structurally, the compound retains the standard L-amino acid backbone (alpha-amino group and alpha-carboxylic acid) in the S absolute configuration, which is critical for ribosomal incorporation. 2.2 Chemical Properties CAS Number: 851960-68-4 IUPAC Name: (S)-2-amino-4-(3-methyldiazirine-3-yl)butanoic acid hydrochloride Solubility: Soluble in water (~10 mg/mL); slightly soluble in DMSO Photoactivation wavelength: ~355–365 nm (UV light) Storage Conditions: –20°C, protected from light; desiccated conditions recommended 2.3 Structural Comparison to L-Methionine The structural similarity between L-Photo-Methionine and the canonical amino acid L-methionine is deliberate and essential to its biological function. L-Methionine possesses a methylthioethyl side chain (-CH2CH2SCH3), whereas L-Photo-Met carries a methyldiazirinylethyl side chain (-CH2CH2C(=N2)CH3). Both side chains are similarly sized, hydrophobic in character, and non-polar. This close structural mimicry is what enables L-Photo-Met to evade the aminoacyl-tRNA synthetase proofreading machinery and be incorporated into proteins during translation in place of methionine. Crucially, the diazirine group — only three atoms in its ring — adds negligible steric bulk compared to many other photocrosslinkers, minimizing perturbation of protein fold and function. 3. Synthesis 3.1 Original Synthesis Route (2005) The synthesis of L-Photo-Methionine was reported by Suchanek, Radzikowska, and Thiele in their landmark 2005 Nature Methods paper. The racemic (DL) form of photo-methionine is synthesized from 4,4′-azipentanal using the Strecker amino acid synthesis — a classical reaction in which an aldehyde is treated with ammonia (or ammonium chloride) and potassium cyanide (KCN) to form an alpha-aminonitrile, which is subsequently hydrolyzed to give the amino acid. The racemic mixture is then resolved to obtain the pure L-enantiomer. Enzymatic resolution of the acetamide derivative is used to separate the enantiomers, exploiting the stereospecificity of appropriate hydrolytic enzymes. The original synthesis yielded racemic photo-methionine at approximately 5%, but the L-enantiomer resolution step was a key challenge. An improved protocol reported a yield of approximately 32% for the synthesis overall — roughly six times greater than the original route — substantially improving the accessibility of the compound for biological research. 3.2 Radiochemical Synthesis In the original characterization studies, a 14C-radiolabeled version of photo-methionine was synthesized to enable autoradiographic detection and to confirm spectroscopic assignments by comparison with known standards. The radiolabeled compound allowed researchers to confirm metabolic incorporation into proteins by tracking radioactivity in protein bands following SDS-PAGE. While radiolabeled versions are not routinely used for modern proteomics (mass spectrometry has largely replaced this approach), they remain valuable for quantitative uptake studies and for confirming the efficiency of metabolic labeling. 4. Mechanism of Action 4.1 Metabolic Incorporation The first and critical step in the use of L-Photo-Methionine in cell biology is its metabolic incorporation into newly synthesized cellular proteins. When cells are grown in medium supplemented with L-Photo-Met (typically at concentrations of 1–4 mM) and depleted of natural methionine, the aminoacyl-tRNA synthetase (MetRS) responsible for charging methionine onto its cognate tRNA (tRNAMet) partially accepts L-Photo-Met as a substrate. Because the structural differences from natural methionine are relatively minor (replacement of -SCH3 with -C(=N2)CH3), the enzyme’s ‘editing’ or proofreading functions are not fully triggered, and the analogue is ligated onto tRNAMet and delivered to the ribosome. At the ribosome, L-Photo-Met is incorporated into growing polypeptide chains wherever a methionine codon (AUG) appears in the mRNA. Since methionine codons are relatively common in proteins (methionine is present on average at ~2.2% frequency in eukaryotic proteomes, and every protein contains at least one methionine as the initiator), L-Photo-Met becomes distributed throughout the newly synthesized portion of the cellular proteome during the labeling window. This results in hundreds to thousands of individual proteins carrying at least one diazirine-modified residue, positioning the photo-crosslinker at or near protein-protein interaction interfaces. 4.2 Photochemical Mechanism: Carbene Generation Upon irradiation with UV light at approximately 355–365 nm, the diazirine ring absorbs photon energy and undergoes electrocyclic photolysis. The mechanistic pathway is now understood to proceed in two sequential steps. First, the diazirine ring opens to generate a diazo intermediate (a species with a =C=N2 moiety), which is more stable and selective than the carbene. The diazo species is particularly reactive toward polar, buried residues that may be inaccessible to conventional chemical crosslinkers. In the second step, the diazo intermediate loses molecular nitrogen (N2) to generate the highly reactive singlet carbene. The singlet carbene produced is an extremely reactive, short-lived species (lifetime on the order of nanoseconds). Importantly, nitrogen gas is the only byproduct released during this process, making the reaction clean and leaving no residual reactive species that could cause off-target damage. The carbene inserts into a wide range of bonds in its immediate molecular environment, including C-H bonds, N-H bonds, and O-H bonds, forming stable covalent adducts. Because the carbene reacts with the nearest available bond — effectively reporting a zero-length crosslink — it captures only genuine, close-contact molecular interactions with high spatial fidelity. 4.3 Reaction Radius and Selectivity A key advantage of diazirine-based crosslinkers is that the carbene reacts within a very short range (essentially zero-length; direct contact required), unlike bifunctional chemical crosslinkers that bridge molecules separated by a spacer arm of defined length. This zero-length characteristic makes L-Photo-Met particularly suitable for mapping protein-protein interfaces with high precision, as only residues that are in direct van der Waals contact with the photo-Met-containing region will be captured. However, the carbene intermediate is non-selective in terms of the chemical bond it attacks — it will react with essentially any C-H, N-H, or O-H bond in its immediate vicinity, regardless of amino acid identity. This lack of chemical selectivity means that mass spectrometric identification of crosslinked residues requires sophisticated bioinformatic tools and can be technically challenging. Recent studies have demonstrated that by tuning irradiation intensity and duration, it is possible to bias the reaction toward the more selective diazo intermediate, which preferentially targets polar, buried residues — providing a new level of structural information inaccessible with other crosslinkers. 4.4 Effect of UV Irradiation on Cell Viability A practical concern with any UV-based crosslinking approach is cell damage. The irradiation conditions required for L-Photo-Met activation (355–365 nm for 1–3 minutes using a high-pressure mercury lamp with a glass filter removing wavelengths below 310 nm) have been empirically validated to be non-damaging to cell viability at these exposure times. Studies have shown that cell viability is maintained after 1–3 minutes of UV exposure under these conditions, with significant cytotoxicity only observed after prolonged exposures of approximately 10 minutes or more. The use of a wavelength above 310 nm is critical: shorter UV wavelengths (UVC, 254 nm, commonly used for sterilization) would cause direct DNA damage and massive cytotoxicity at the doses required for efficient crosslinking. 5. Experimental Protocols and Practical Considerations 5.1 General Metabolic Labeling Protocol The following outlines a generalized protocol for metabolic labeling of mammalian cells with L-Photo-Methionine for photoaffinity crosslinking of PPIs: • Culture cells to approximately 60–80% confluency in normal complete medium. • Remove standard culture medium and wash cells once with PBS to remove residual methionine. • Replace with methionine-free, leucine-free medium (if co-labeling with photo-leucine) without phenol red, which can absorb UV light. • Add L-Photo-Met to a final concentration of approximately 1.7–4 mM (titrate for optimal incorporation vs. cytotoxicity). • Incubate cells for 18–24 hours to allow sufficient metabolic incorporation into newly synthesized proteins. • Wash cells twice with ice-cold PBS to remove unincorporated photo-Met. • Place culture dishes on ice and irradiate with a 200 W high-pressure mercury lamp (or equivalent UV source) fitted with a glass filter to remove wavelengths below 310 nm for 1–3 minutes. • Lyse cells immediately on ice using an appropriate lysis buffer. • Analyze crosslinked products by SDS-PAGE and western blotting, immunoprecipitation, or mass spectrometry-based proteomics. 5.2 Combination with Photo-Leucine L-Photo-Methionine is frequently used in combination with its structural analogue L-Photo-Leucine (a diazirine-containing leucine analogue) in the same experiment. Since both are incorporated by the cellular translational machinery using their respective aminoacyl-tRNA synthetases, co-supplementing cells with both analogues dramatically increases the probability of photo-crosslinker incorporation at protein-protein interfaces (since any surface methionine or leucine residue can carry the reactive group). In the original Suchanek et al. protocol, cells were supplemented with 4 mM photo-leucine, 1.7 mM photo-methionine, and 4 mM photo-isoleucine for 22 hours. This multi-analogue approach is now standard in many labs for comprehensive capture of protein complexes. 5.3 Solid-Phase Peptide Synthesis Applications The Fmoc-protected derivative of L-Photo-Met is used in SPPS to create synthetic peptide probes incorporating the photo-crosslinker at a specific, defined position. This approach is employed when researchers need to study PPIs at known binding interfaces, probe the binding pocket of a protein target, or generate photo-reactive versions of known bioactive peptides. Fmoc-L-Photo-Met is compatible with standard Fmoc SPPS protocols using HATU/DIPEA or similar coupling conditions, though care must be taken to minimize exposure to light during synthesis and to store the resin-bound intermediate protected from UV. 5.4 Downstream Analysis: Western Blotting and Mass Spectrometry Following UV-induced crosslinking, the resulting covalent protein complexes must be analyzed. For simple detection of crosslinked species, standard SDS-PAGE under denaturing conditions followed by western blotting using antibodies against proteins of interest is sufficient. Crosslinked dimers, trimers, and higher-order oligomers appear as higher-molecular-weight bands compared to non-crosslinked controls. For unbiased identification of crosslinked proteins and the precise crosslinked residues, high-resolution mass spectrometry (MS and MS/MS) is the gold standard. Crosslinked peptides are identified by their characteristic mass shifts. Dedicated bioinformatic pipelines (e.g., pLink, StavroX, MeroX, and others) are used to search databases for crosslinked peptide pairs and map the spatial constraints onto protein structures. The zero-length nature of the L-Photo-Met crosslink provides stringent distance restraints (Cα-Cα distance≤10-12 Å) that can be used in integrative structural biology workflows to validate or refine protein complex models. 5.5 Storage and Handling L-Photo-Methionine must be stored at –20°C in a dark environment (e.g., amber vials or foil-wrapped containers) to prevent premature photolysis of the diazirine ring by ambient light. The lyophilized powder is relatively stable under these conditions. Working solutions should be prepared fresh and kept on ice and in the dark until use. Prolonged exposure to fluorescent laboratory lighting can degrade the diazirine, leading to reduced crosslinking efficiency 6. Applications 6.1 Mapping Protein-Protein Interactions in Living Cells The flagship application of L-Photo-Methionine is the direct identification and characterization of protein-protein interactions in their native cellular environment. Unlike co-immunoprecipitation, which relies on antibody affinity capture under potentially disruptive lysis conditions, L-Photo-Met crosslinking occurs inside intact, living cells under physiological conditions. This is a critical distinction because many biologically relevant PPIs — particularly transient signaling interactions, receptor-effector contacts, and stoichiometrically minor complexes — are disrupted by the dilution, detergents, and non-native salt conditions encountered during cell lysis. The original landmark application by Suchanek et al. demonstrated this power by identifying a previously unknown direct protein-protein interaction involving membrane proteins regulating cholesterol homeostasis: SCAP (SREBP cleavage-activating protein), Insig-1, and SREBP (sterol regulatory element-binding protein) were shown to form a direct complex. Additionally, the researchers employed photo-Met to demonstrate a direct interaction of the progesterone receptor membrane component 1 (PGRMC1) with a novel binding partner, a finding that would have been difficult or impossible using non-crosslinking methods. 6.2 Membrane Protein Complexes Membrane proteins represent a particularly challenging class of targets for PPI studies because they are embedded in lipid bilayers, which are disrupted by most standard detergent-based lysis protocols. L-Photo-Methionine is exceptionally well-suited to membrane protein complex studies because crosslinking occurs while the membrane is intact, prior to any disruption. The covalent bond formed during UV irradiation holds the complex together even through subsequent detergent solubilization, affinity purification, and denaturing electrophoresis. This has enabled the characterization of multiple receptor complexes, ion channel assemblies, and transporter oligomers that were previously intractable. 6.3 Post-Translational Modification (PTM)-Dependent Interactions Many protein-protein interactions are regulated by post-translational modifications such as phosphorylation, ubiquitination, acetylation, and methylation. Because these modifications are transient and substoichiometric, capturing PTM-dependent PPIs is particularly challenging. L-Photo-Methionine can be combined with expressed protein ligation (EPL) or semi-synthesis approaches to introduce both a specific PTM (e.g., a phosphoserine) and the photo-crosslinker into the same protein molecule, allowing the researcher to ask whether the interaction they observe is strictly dependent on the modification. A notable example is the use of Smad2-MH2 constructs bearing both a phosphoserine (pSer) and a photo-Met residue to demonstrate that MH2-MH2 trimerization is strictly dependent on receptor-mediated serine phosphorylation. This type of experiment, demonstrating a transient, PTM-dependent interaction, is uniquely enabled by L-Photo-Met technology. 6.4 Target Identification in Drug Discovery One of the most exciting modern applications of photo-reactive amino acids, including L-Photo-Met, is in target identification (target ID) and validation in drug discovery. A major challenge in phenotypic drug discovery — where a compound is identified by its cellular effect without knowledge of its molecular target — is subsequently identifying which protein(s) the compound engages. Chemical proteomics approaches using photoaffinity probes are a powerful solution: a drug candidate is modified to incorporate a diazirine group and optionally a clickable handle (alkyne or azide) for downstream enrichment via bioorthogonal click chemistry. The probe is then added to living cells, allowed to bind its protein target(s), and covalently captured by UV irradiation. Enrichment and MS identification then reveal the target interactome. L-Photo-Met has been used as a building block in such multifunctional probes, where its structure contributes the photocrosslinkable diazirine in a scaffold that mimics a natural methionine in drug-like molecules derived from S-adenosylmethionine (SAM) pathways, methyltransferase inhibitors, or other methionine-utilizing biochemical contexts. 6.5 Structural Proteomics and Protein Structure Determination Photo-crosslinking with L-Photo-Met generates distance restraints that can be used in structural proteomics. When MS identifies a crosslinked peptide pair, it establishes that the two crosslinked residues must be within direct contact distance in the native protein complex. This information provides experimental constraints that can be used in integrative structural modeling to build or validate models of protein complexes for which high-resolution structural data (X-ray crystallography, cryo-EM) are unavailable. A published example involves using a protein nanoprobe incorporating photo-Met to map the homodimerization interface of the 14-3-3ζ protein, where high-resolution MS was used to determine both the crosslinked residues and the reaction radius, yielding structural information consistent with the known crystal structure. 6.6 Proteome Profiling with Photo-Amino Acids Beyond the study of individual PPIs, L-Photo-Met has been applied to global proteome-wide interaction profiling. When cells are labeled with photo-Met and the entire crosslinked proteome is subjected to MS-based analysis, interaction networks can be mapped at the systems level. This approach — sometimes called global photo-crosslinking mass spectrometry (GP-XL-MS) — complements affinity purification-MS approaches by capturing transient and low-affinity interactions that are routinely lost in AP-MS workflows. Recent advances in crosslink-enrichment strategies, high-sensitivity MS instrumentation, and bioinformatic pipeline development have substantially increased the depth and coverage achievable in such experiments. 7. Comparison with Other Photocrosslinking Strategies 7.1 Overview of Major Photocrosslinker Classes Three major classes of photocrosslinker are in widespread use: diazirines (DAs), benzophenones (BPs) and aryl azides (AAs). Each has distinct photochemical properties, reactive intermediates, activation wavelengths, and biological suitability profiles. The following is a head-to-head comparison of L-Photo-Met (as a representative diazirine) with BP and AA crosslinkers: L-Photo-Met (Diazirine) UV wavelength: 355–365 nm Reactive intermediate: Carbene / diazo Crosslink size: Zero-length (very small) Cell permeability: High (metabolic incorporation) Protein perturbation: Minimal (methionine mimic) Selectivity: Non-selective C-H, N-H, O-H insertions Background labeling: Low-moderate Living cell suitability: Excellent Benzophenone UV wavelength: 350–365 nm Reactive intermediate: Triplet diradical Crosslink size: Moderate Cell permeability: Probe-dependent Protein perturbation: Moderate (bulky group) Selectivity: Preference for C-H near heteroatoms Background labeling: Low Living cell suitability: Good Aryl Azide UV wavelength: 250–280 nm Reactive intermediate: Nitrene Crosslink size: Zero-length Cell permeability: Probe-dependent Protein perturbation: Moderate Selectivity: Nitrene ring expansion possible Background labeling: Moderate Living cell suitability: Limited (short UV) 7.2 Advantages of L-Photo-Methionine Over Conventional Photocrosslinkers The principal advantage of L-Photo-Met over conventional exogenous photocrosslinkers (whether diazirine, benzophenone, or aryl azide based) is its mechanism of delivery: metabolic incorporation. Conventional photoaffinity probes must be chemically conjugated to a molecule of interest (a drug, a peptide, a lipid) and then added exogenously to cells or cell lysates. This requires synthetic derivatization of the probe molecule and relies on the probe being cell-permeable, reaching its target at sufficient concentration, and not being significantly metabolized or sequestered before reaching its target. L-Photo-Met sidesteps all of these concerns by exploiting the cell’s own translational machinery. The photo-crosslinker is distributed by the ribosome throughout all newly synthesized proteins in the cells, without any requirement for chemical modification of a specific probe molecule. This makes it an unbiased, proteome-wide crosslinking approach rather than a targeted one. It is also suitable for studying PPIs of endogenous proteins in their native stoichiometry and subcellular localization — a significant advantage over overexpression-based approaches. 7.3 L-Photo-Met vs. Genetic Code Expansion Approaches An alternative strategy for incorporating photoreactive amino acids into proteins is genetic code expansion (GCE), in which an orthogonal aminoacyl-tRNA synthetase/tRNA pair is engineered to incorporate an unnatural amino acid in response to a stop codon (usually the amber UAG codon) at a defined site in the protein of interest. GCE enables precise, site-specific incorporation at a single, pre-determined position in a specific protein, offering extremely high resolution for mapping a specific binding interface. However, GCE requires extensive molecular biology work (plasmid construction, stable cell line generation), works only in the target protein (not proteome-wide), and typically requires suppression of a stop codon which can reduce expression levels. L-Photo-Met, by contrast, requires no genetic manipulation and labels the entire newly synthesized proteome, making it ideal for unbiased discovery of novel interactions. The two approaches are thus complementary: L-Photo-Met for unbiased, proteome-wide PPI discovery, and GCE for high-resolution site-specific validation of specific interactions. 8. Limitations and Challenges 8.1 Non-Site-Specific Incorporation The most significant limitation of L-Photo-Met is that it cannot be incorporated at a single defined position in a specific protein of interest. It is incorporated at every methionine position in every newly synthesized protein. This means that in a given experiment, crosslinks can arise from any methionine-bearing region of any protein, not specifically from an interface of interest. For global interaction studies this is a feature, but for precise structural questions requiring a specific crosslink at a specific residue, the Fmoc-SPPS approach or GCE must be used instead. 8.2 Requirement for Methionine Starvation To achieve efficient incorporation, cells must be methionine-starved for many hours (typically 18–24 h). This starvation is itself a significant cellular stress and can activate methionine-sensitive stress response pathways, potentially altering the protein interaction landscape being studied. Extended methionine starvation can also impair cell growth, alter translation rates, and confound results. Researchers must carefully control for these effects, for example by comparing cells grown in methionine-free medium with and without photo-Met, and limiting the starvation period where possible. 8.3 Competition with Natural Methionine Even under methionine starvation conditions, intracellular methionine pools may not be fully depleted, particularly from stored/recycled methionine from protein turnover. Residual natural methionine competes with L-Photo-Met for incorporation, reducing labeling efficiency. Higher concentrations of L-Photo-Met can be used to shift the competition in favor of the analogue, but very high concentrations may themselves be cytotoxic or alter translation efficiency. 8.4 Identification of Crosslinked Residues by MS While MS is the gold standard for identifying crosslinked proteins and residues, the analysis is technically demanding. Diazirine crosslinks create mass shifts that must be matched against theoretical databases, and the many possible bond insertion sites (C-H, N-H, O-H at any residue) mean that the search space is very large. The efficiency of crosslink formation at any given contact site is typically low (1–10%), meaning that crosslinked peptides are low-abundance species in a complex mixture. Enrichment strategies and high-sensitivity MS are required. Dedicated software tools for crosslink identification (pLink, StavroX, MeroX, Scout, and others) are available but require expertise to use effectively. 8.5 Photodamage and Background Crosslinking While UV irradiation at 365 nm is relatively gentle, prolonged or high-intensity irradiation can cause oxidative damage to proteins and nucleic acids, generating false crosslinks. The diazirine carbene, once generated, will react with the nearest bond — which may not always be an interacting protein, but could be another region of the same protein (intramolecular crosslink), a lipid, a nucleic acid, or simply water (hydrolysis). High background crosslinking reduces the signal-to-noise ratio in PPI studies. The diazo intermediate pathway, which precedes carbene formation, offers slightly better selectivity and can be favored by reducing irradiation intensity. 9. Related Derivatives and Analogues 9.1 Fmoc-L-Photo-Methionine Fmoc-L-Photo-Methionine (CAS #: 945859-89-2) is the N-terminally protected derivative designed for solid-phase peptide synthesis. The 9-fluorenylmethoxycarbonyl (Fmoc) protecting group is stable under standard peptide synthesis conditions (piperidine deprotection, TFA cleavage) and is routinely used in Fmoc SPPS. This derivative allows synthetic chemists to prepare peptides containing a photo-Met residue at any desired position, enabling highly targeted photoaffinity labeling studies of peptide-protein interactions, receptor binding, and enzyme-substrate contacts. 9.2 L-Photo-Leucine L-Photo-Leucine (CAS #: 851960-91-3) is the leucine analogue bearing a diazirine ring in place of the isobutyl side chain, designed and synthesized alongside L-Photo-Met by the same research group. Like photo-Met, it is metabolically incorporated by the cellular translational machinery and activatable at ~365 nm. Because leucine is more abundant than methionine in most proteomes (~9% vs. ~2.2%), photo-Leu typically achieves higher overall labeling density. The two analogues are frequently used together in the same experiment to maximize coverage. 9.3 L-Photo-Isoleucine L-Photo-Isoleucine is a further analogue designed to mimic isoleucine. It is co-supplemented in some protocols alongside photo-Met and photo-Leu for comprehensive labeling. Its use is less common than the Met and Leu analogues due to more challenging synthesis and somewhat lower incorporation efficiency. 9.4 Bifunctional Probes Incorporating Photo-Met A growing class of multifunctional chemical biology probes incorporate the photo-Met diazirine moiety as one of several functional groups in a single molecule designed for drug target identification. These probes typically combine: (1) a pharmacophore that binds the target protein non-covalently, (2) the diazirine group for UV-induced covalent capture, and (3) a bioorthogonal handle (terminal alkyne or azide) for click chemistry-mediated enrichment and detection. Such probes have been used to identify the cellular targets of a variety of drug candidates and natural products in unbiased chemoproteomic screens. 10. Recent Advances and Future Directions 10.1 Mechanistic Understanding of Diazirine Photochemistry A 2024 study published in Nature Communications provided important new mechanistic insight into alkyl diazirine photo-crosslinking, including that used by photo-Met. Using a systematic modulation of light intensity and irradiation time, the researchers quantitatively dissected the diazirine photolysis pathway and confirmed a sequential two-step mechanism: diazirine → diazo intermediate → carbene. Critically, they showed that the diazo intermediate preferentially cross-links to polar, buried residues that are inaccessible to conventional chemical crosslinkers, and that by tuning irradiation conditions, researchers can bias the crosslinking reaction toward these more informative, structurally discriminating products. This new understanding has practical implications for optimizing L-Photo-Met experiments. 10.2 Integration with Click Chemistry Modern versions of photo-Met-based approaches increasingly combine metabolic labeling with bioorthogonal click chemistry. By co-supplementing cells with azidohomoalanine (AHA, a methionine analogue bearing an azide group) alongside photo-Met in sequential or simultaneous labeling, researchers can enrich newly synthesized, photo-Met-containing proteins using strain-promoted or copper-catalyzed azide-alkyne cycloaddition. This combination of metabolic labeling, photo-crosslinking, and bioorthogonal enrichment greatly reduces background and increases sensitivity for identifying rare or transient interaction partners. 10.3 Applications in Structural Proteomics Pipelines The integration of in-cell photo-crosslinking with integrative structural modeling is a rapidly maturing field. Photo-Met-derived crosslinking restraints, when combined with cryo-EM maps, AlphaFold2 structural predictions, and other experimental data types in platforms such as IMP (Integrative Modeling Platform), enable the construction of high-confidence models of large macromolecular assemblies. As the cost of MS decreases and the sensitivity of crosslink identification improves, photo-Met crosslinking is expected to become a routine component of integrative structural proteomics pipelines. 10.4 Expansion to Non-Mammalian Systems While the majority of L-Photo-Met applications have been demonstrated in mammalian cell lines, there is growing interest in its use in other biological systems. Metabolic incorporation has been reported in bacterial systems (particularly in E. coli strains with relaxed methionine aminoacyl-tRNA synthetase editing), in yeast, and in primary cell cultures. Expansion to model organisms such as Drosophila and C. elegans is an active area of development, as this would allow photo-crosslinking studies in the context of a whole organism. 10.5 Future Outlook L-Photo-Methionine and the broader field of photo-reactive non-canonical amino acids continue to evolve rapidly. Key future directions include: the development of genetically encodable versions that can be controlled with greater spatial and temporal precision; the combination with proximity ligation approaches such as TurboID or APEX2 for orthogonal validation; the use of multi-wavelength photoactivatable systems that allow sequential activation of different crosslinkers within the same experiment; and the coupling of photo-crosslinking with single-cell proteomics approaches to interrogate PPI heterogeneity at the level of individual cells. 11. Conclusions L-Photo-Methionine (CAS #: 851960-68-4) is a pioneering reagent that has made lasting contributions to the chemical biology toolkit for studying protein-protein interactions. Its design — a diazirine-bearing structural mimic of L-methionine that is recognized and incorporated by the cellular translational machinery — represented a conceptually elegant solution to the challenge of introducing photo-crosslinking capability into the cellular proteome without genetic manipulation. The two decades since its first description have seen it become a well-established tool in cell biology, membrane biochemistry, structural proteomics, and drug discovery research. Its defining advantages are its ability to study PPIs in their native cellular context, its compatibility with living cells under physiological conditions, and the precision of the zero-length diazirine crosslink. These properties, combined with the development of companion analogues (photo-Leu, photo-Ile), Fmoc-protected derivatives for SPPS, and integration with modern MS, click chemistry, and structural modeling tools, ensure that L-Photo-Met remains a highly relevant and powerful research reagent. Challenges remain, particularly around the non-specific distribution of labeling across the proteome, the technical demands of MS-based crosslink identification, and the practical handling restrictions associated with the self-reactive diazirine group. Nevertheless, ongoing methodological advances — including improved understanding of the two-step diazo/carbene mechanism, coupling with click chemistry enrichment, and integration into integrative structural proteomics pipelines — continue to expand the scope and resolution of what can be achieved with this remarkable chemical tool. As the field of chemical biology moves toward ever more precise, quantitative, and proteome-wide interrogation of molecular interactions in living systems, L-Photo-Methionine will undoubtedly continue to occupy an important role at the frontier of discovery. References 1. Suchanek M, Radzikowska A, Thiele C. Photo-leucine and photo-methionine allow identification of protein-protein interactions in living cells. Nat Methods. 2005;2(4):261-268. doi:10.1038/nmeth752 2. Dubinsky L, Krom BP, Meijler MM. Diazirine based photoaffinity labeling. Bioorg Med Chem. 2012;20(2):554-570. doi:10.1016/j.bmc.2011.06.066 3. Hashimoto M, Hatanaka Y. Photo-affinity labeling (PAL) in chemical proteomics: a handy tool to investigate protein-protein interactions (PPIs). Proteome Sci. 2017;15(1):13. doi:10.1186/s12953-017-0123-3 4. Halloran MW, Lumb JP. Recent Applications of Diazirines in Chemical Proteomics. Chem Eur J. 2019;25(20):4885-4898. 5. Das J. Aliphatic diazirines as photoaffinity probes for proteins: recent developments. Chem Rev. 2011;111(7):4405-4417. doi:10.1021/cr1002722 6. Brunner J. New photolabeling and crosslinking methods. Annu Rev Biochem. 1993;62:483-514. 7. Smith E, Collins I. Photoaffinity labeling in target- and binding-site identification. Future Med Chem. 2015;7(2):159-183. 8. Murale DP, Hong SC, Yun WS, Lee JS. Photo-affinity labeling (PAL) in chemical proteomics. Proteome Sci. 2017;15:14. 9. Yang T et al. Dissecting diazirine photo-reaction mechanism for protein residue-specific cross-linking and distance mapping. Nat Commun. 2024;15:6093. doi:10.1038/s41467-024-50315-y 10. Walrant A et al. Photoaffinity labeling coupled to MS to identify peptide biological partners. Mass Spectrometry Reviews. 2024. doi:10.1002/mas.21880 11. Benns HJ et al. Photoaffinity labelling strategies for mapping the small molecule-protein interactome. Curr Opin Chem Biol. 2021;60:112-121. |
|
H-Photo-Met-OH
For Research & Development use only. Not for testing and/or use on humans.